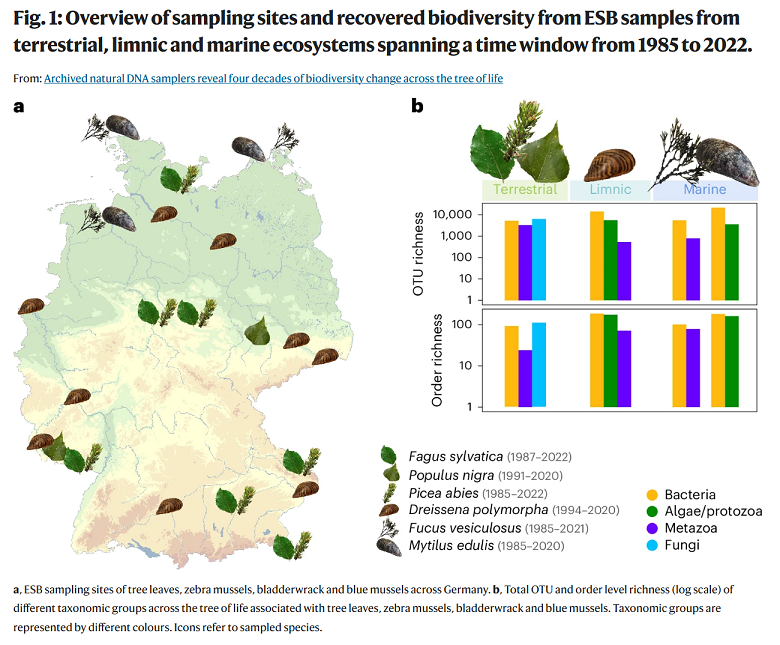
Archived natural DNA samplers reveal four decades of biodiversity change across the tree of life
Junk, Isabelle; Hans, Julian; Perez-Lamarque, Benoît; Stothut, Manuel; Weber, Sven; Gold, Elisabeth; Schubert, Caroline; Schumacher, Alice; Schmitt, Nina; Melcher, Anja; Paulus, Martin; Klein, Roland; Teubner, Diana; Koschorreck, Jan; Kennedy, Susan; Morlon, Hélène; Krehenwinkel, Henrik
Nature Ecology & Evolution
Detecting the imprints of global environmental change on biological communities is a paramount task for ecological research. But a lack of standardized long-term biomonitoring data prevents a deeper understanding of biodiversity change in the Anthropocene. Novel sources of data for analysing biodiversity change across time and space are urgently needed. By metabarcoding highly standardized biota samples from a long-term pollution monitoring archive in Germany, we here analyse four decades of community diversity for tens of thousands of species across the tree of life.
The archived samples—tree leaves, marine macroalgae, and marine and limnic mussels—represent natural community DNA samplers, preserving a taxonomically diverse imprint of their associated biodiversity at the time of collection. We find no evidence for universal diversity declines at the local scale. Instead, a gradual compositional turnover emerges as a universal pattern of temporal biodiversity change in Germany’s terrestrial and aquatic ecosystems.
This turnover results in biotic homogenization in most terrestrial and marine communities. Limnic communities, in contrast, rather differentiate across space, probably due to the immigration of different invasive species into different sites. Our study highlights the immense promise of alternative sample sources to provide standardized time series data of biodiversity change in the Anthropocene.
doi.org/10.1038/s41559-025-02812-6